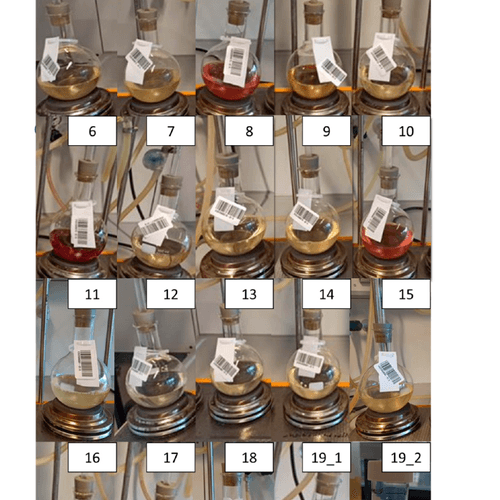
Grain and Seed Phenotyping
Built for Research
High-throughput, non-destructive phenotyping for grains and seeds. Quantify morphology, L*a*b* color, defects, and purity with NIST-traceable measurements. Export bioinformatics-ready CSV/Parquet outputs compatible with R and Python pipelines.

Validated By
University and research labs worldwide
Standardized
Protocols across operators and seasons
Publication-Ready Data
Outputs (CSV + images + metadata)
Research-grade
Consistency across sites and time
Trusted by Leading Institutions
High-throughput phenotyping programs at agricultural universities, research centers, and breeding labs rely on Vibe for reproducible, non-destructive grain and seed imaging with genotype-to-phenotype workflows that stay consistent across operators, seasons, and sites.
Peer-reviewed publications using Vibe
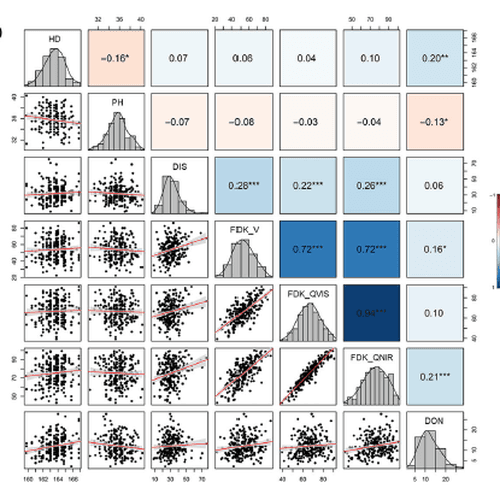
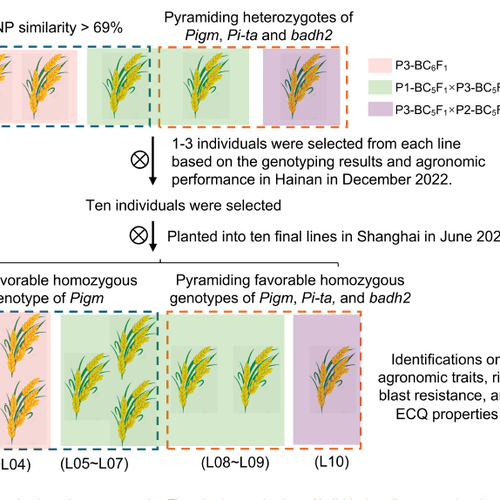
Peer-reviewed evidence from breeding programs and grain quality labs using Vibe for high-throughput phenotyping (HTP), morphological profiling, and non-destructive seed analysis. Browse the publications library for methods, citations, and genotype-to-phenotype research.
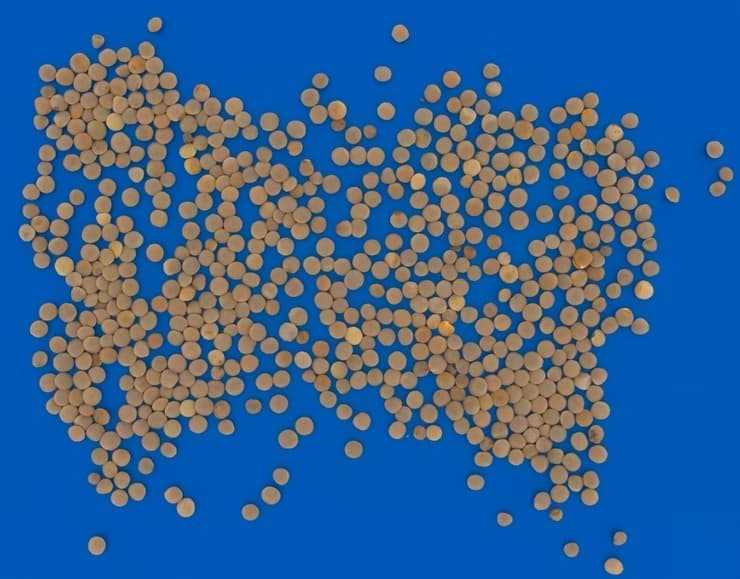
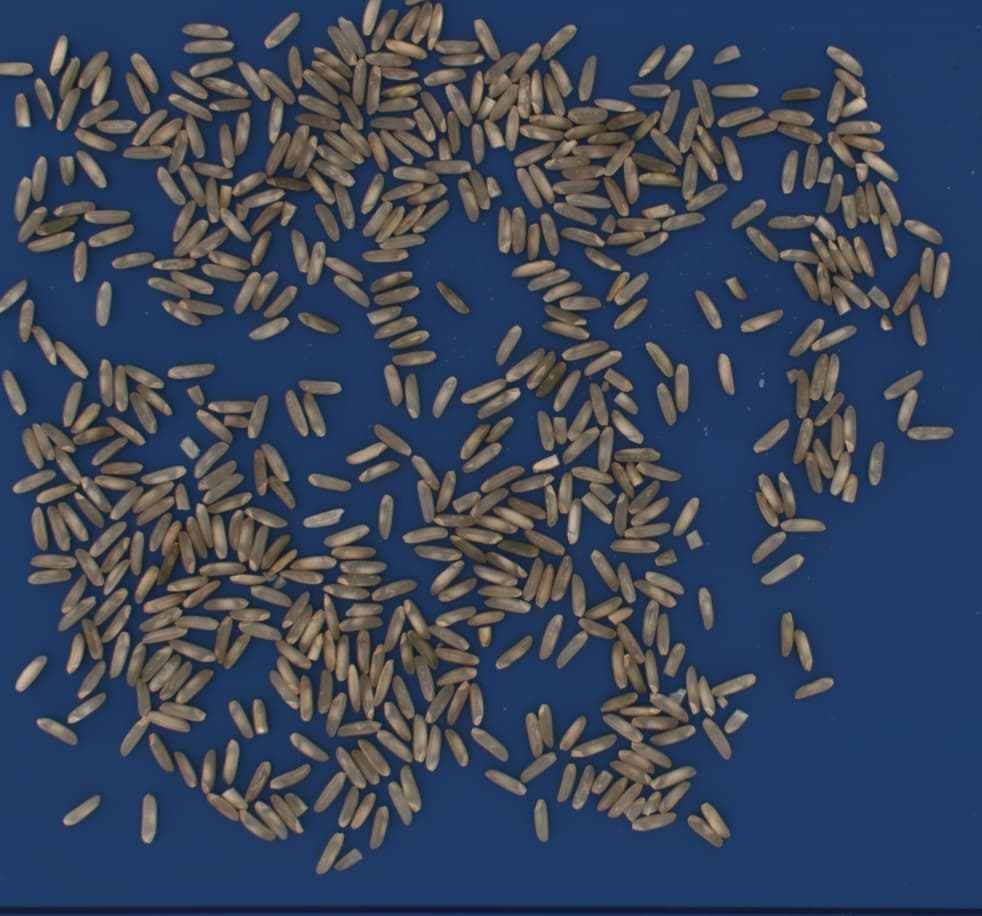
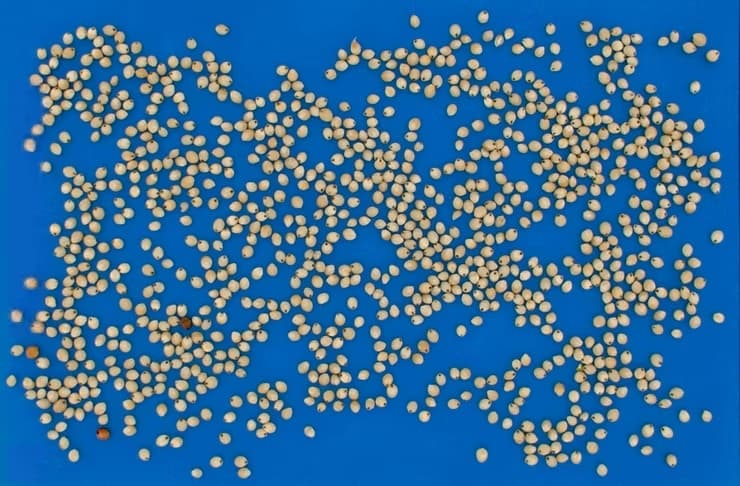
Crop types commonly analyzed in research labs
From morphological profiling of wheat endosperm to vitreousness scoring in durum - workflows handle varieties, mixtures, and custom label definitions for any crop.
Key outputs
Per-kernel morphological profiling, size distributions, broken percent, defect indicators, purity metrics, and NIST-traceable L*a*b* color features - all exportable for downstream bioinformatics and R/Python analysis.
Lab workflows and protocols
Practical workflows for purity testing, grading, defect detection, morphology, and more. Each workflow shows what to measure, how to run it, and which outputs to export. Explore the workflow library for additional examples.
Built for publishable, repeatable grain and seed research
Vibe helps research teams run high-throughput, non-destructive phenotyping with NIST-traceable measurements comparable across operators, seasons, and sites. This hub collects the evidence: peer-reviewed publications, bioinformatics-ready peer-reviewed evidence, and validated lab workflows.
Common questions from research teams
Short answers focused on repeatability, workflows, and exports.
Vibe is a high-throughput, non-destructive imaging platform for reproducible grain and seed phenotyping - built so research labs get NIST-traceable measurements every run.
Per-kernel morphological profiling (length, width, area, shape), NIST-traceable L*a*b* color, defect indicators, purity, broken percent, and batch comparisons - all exportable as CSV/Parquet for R and Python pipelines.
Yes. The QM3i Analyzer uses optical imaging - samples are not altered, so the same material can be used for downstream moisture, milling, or genetic analysis.
Yes. Exports include structured CSV/Parquet tables with run metadata so downstream analysis is fully reproducible and figures can be regenerated without manual steps.
Yes. Under 1 minute per 40 g sample means hundreds of accessions can be processed in a single session, making it practical for large breeding trials and genotype-to-phenotype studies.
Yes. Queue multiple samples with barcode scanning for continuous, unattended processing — designed for breeding programs that need to phenotype hundreds of lines per session.
Most start with a short demo, confirm the protocol and outputs match their study needs, then run a limited-scope pilot before expanding to more crop types and workflows.