Improving panicle blast resistance and fragrance in a high-quality japonica rice variety through breeding
Shanghai Academy of Agricultural Sciences
Partner institutions
Shanghai Academy of Agricultural Sciences
Shanghai, China
Rice breeding and quality studies; applied screening workflows with Vibe images.
How Vibe analyzers were used
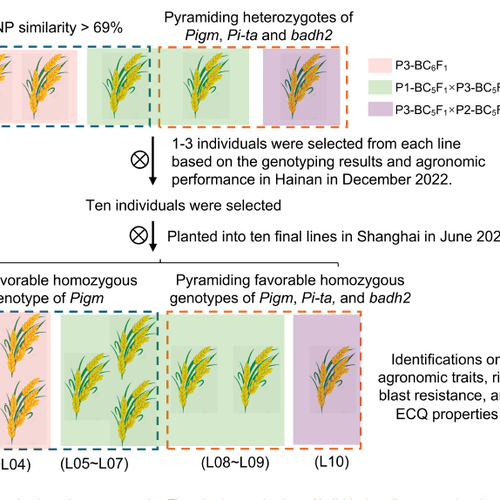
Marker-assisted backcrossing introgressed blast-resistance genes (Pigm, Pi-ta) and the aroma gene badh2 into Huruan1212, producing lines with improved panicle blast resistance and fragrance while maintaining high eating and cooking quality.
Study overview
Huruan1212 (HR1212) is a high-quality japonica rice variety but is susceptible to panicle blast and lacks fragrance. The study used marker-assisted selection and resequencing-based background selection to introduce Pigm, Pi-ta, and badh2. Three introgressed lines were selected, showing enhanced resistance to panicle blast, improved yield relative to HR1212, and (for the pyramided line) strong aroma, while maintaining eating and cooking quality comparable to HR1212.
Topics and keywords
Bibliographic details
Year: 2025
Publication: Frontiers in Plant Science
DOI: 10.3389/fpls.2024.1507827
Institutions: Shanghai Academy of Agricultural Sciences


